Peering through the tissues #1
A primer on how cells organize and how scientists are using spatial omics to dig deeper

Imagine you are Robert hooke living in the late 1600s with the brand new state-of-the-art compound microscope and you take frequent walks in the gardens to pick your samples just out of curiosity; to see what exists at much smaller scales and you happen to observe a range of diverse organisms and you start out to wonder why this tiny insect has such complex structures that are not visible to the naked eye but they exists you wonder what these segments do. But you are still limited to peer deeper. your curiosity persists but the technology doesn't allow to examine deeper. you have to wait for another hundred or so years for a description of what you see. You see compartmentalisation of certain specimens and vague descriptions to the best of current understanding.
A hundred years later, you are Xavier Bichat grinding through your studies in anatomy and surgery in Lyon. through your keen and astute observations, you describe what a tissue is, and you are successfully able to distinguish 21 elementary types of tissues; but you are still unable to ascertain exact functions and what they compose of; You have to probably wait a few more decades so you would understand the organisational units and how the structure and physiology of living things and how these form the lego blocks of sorts to the tissue and ultimately the organ. But you are still in awe about the complexity and function of the individual cells in this.
Welcome to the 1970s the methods are much more advanced than the hooke microscope or the observational studies you as in your previous imaginary avatars have had access to. Now you see the scientists trying to quantify messenger RNA of the genes in tissues and cells. They are using altered forms of in-situ hybridisation to achieve this; one such method is Radioactive-ISH. Then, the path to further studies such as visualizing transcripts of specific genes has become possible. Furthermore, a decade or so later it was possible via colourimetric ISH to visualize and understand the transcriptional processes in 3 dimensions.
It’s the 80s and 90’s now DNA sequencing become slowly but steadily the state-of-the-art technologies. For the first time since Hooke’s observations, we can perform experiments that visualize the expression of unknown genes. The foundation for the latest generation spatial assays and transcriptional analysis of tissues has been laid.
The motivation behind such an arduous and lengthy journey has been identifying genes with restricted patterns in individual cells. These genes may participate in development and aid in the identification of novel cell types and markers that are not distinguishable by tissue morphology. By this era, the convergence of technologies including the availability of cheaper and higher throughput sequencing technologies and more powerful computing infrastructure has made spatial transcriptomics from a niche field to a much more regular and mainstream affair.
Spatial omics technologies: A walk-through of what tools are available
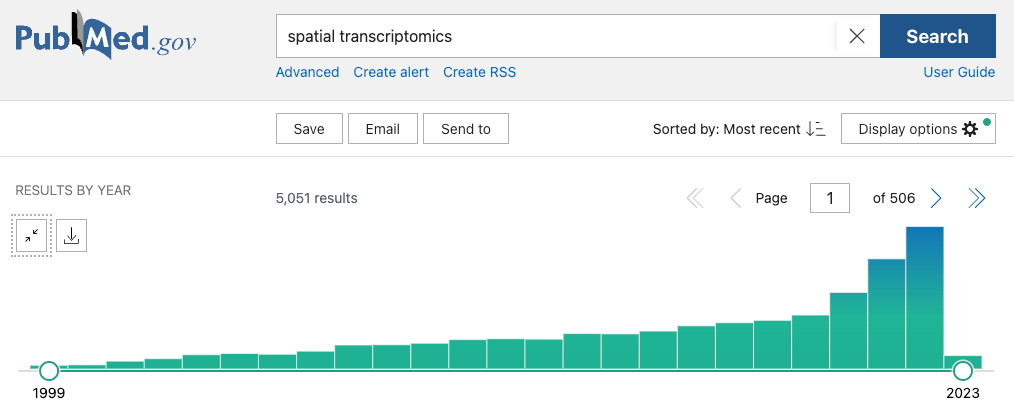
The recent advancements in the development of spatial omics technologies and the breadth of choice of these technologies might be a bit intimidating for a newbie researcher to approach and integrate them into their workflow. There was a similar case for me in a previous lab where we wanted to implement scSEQ technology to understand the expression patterns of Epstein Barr virus in patients suffering from PTLD and we had so many methods to screen through. So let us break down what is available for you.
Broadly speaking, the technologies vary in
Resolution i.e., how accurately can we distinguish the cells etc
coverage i.e., what tissues can be covered
scale and throughput ie number of samples that can be processed and how fast can it be done
multiplexing capacity ie how many genes or mRNA can be measured simultaneously
With this in mind, a researcher can approach their question of choice based on profiling methods:
Targeted, multiplexed probe or antibody-based: ideal if you want to study a certain surface marker for a cell within a tissue or if you have a panel of targets that you are interested in; this could be oncogenes within lung cancer tissue for example.
transcriptome-wide or NGS-based approaches if you are interested in a specific condition and you want to profile the global expression of cells within tissues.
The following table is a rough estimate of the resolution required when using spatial transcriptomics in your projects.
A gentle intro to the technologies at hand
You may ask, Hey dude why should I bother? or There are so many I will ask a postdoc to sort my experiment out. Seeking guidance is probably the best way forward but I want to try to explain to myself and in the process probably teach you what is available. I will certainly do a deep dive of sorts into these available technologies based on their popularity
You have a target in mind: This is likely your way!
It’s often the case that initial exploratory analysis via traditional/conventional experimentation leads to an initial hypothesis to be tested using targeted spatial omics. In these cases, Antibodies and RNA probes are used as your detection methods. One of the interesting methods that led to the development of multiplexed assays in the targeted space is from the sequencing space; where in one can use antibodies that contain cleavable linkers for sequential imaging. This was one of a project heavily focussed on Ulf Landegrens's lab at IGP Uppsala university that eventually evolved into Cartana now acquired by 10x Genomics when i was a research trainee. By using barcoding schemes researchers can profile more than 50 cellular entities in a single slide section.

Imaging-based approaches
Currently, there are two go-to methods when it comes to imaging-based approaches that have matured and you have a choice of companies providing kits etc.,
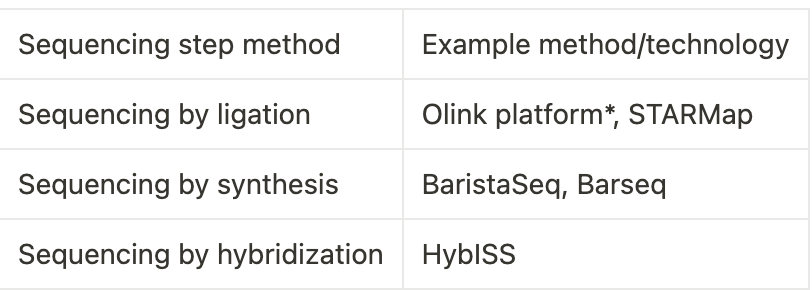
In-situ sequencing methods (ISS) leverage the age-old rolling-circle amplification to read out the sequences of transcripts within the tissue. To achieve this RNA is reverse transcribed, consequently amplified and sequenced. The probes used are targetted for reverse transcription and followed by sequencing by ligation. Other variations of this is by altering the probes to fit one of the following sequencing methods sequencing by synthesis, or sequencing by hybridization. ISS has also been used for untargeted profiling with a limitation that there might be optical overcrowding (interpret this as overlapping of the amplicons and general indistinguishability between different transcripts) which leads to lower sensitivity. One exception to this is ExSeq which uses expansion microscopy to perform untargeted ISS in tissues.

In-situ Hybridization methods are the second category of imaging-based approach. Here, the target sequence is detected by the hybridization of a complementary fluorescent probe. Initially, this limited number of transcripts can be probed, but subsequent innovations led to higher multiplexing. One established example is MERFISH which uses serial imaging post-hybridisation rounds to identify the presence or absence of transcripts in the cells; MERFISH has been used over various applications to profile transcript locations within cells and tissues. By using a combination of colours seqFISH has developed an expanded barcode palette of 60 pseudocolours divided in three fluorescent channels. This method can generate large maps of tissues and organ sections. The methods are constantly being improved and if Moore’s law can be applied to biology we will see a further jump in the total number of genes detected at a sub-cellular resolution.
So how do these work and what’s the role of Bioinformatics here you might ask…
Essentially after getting the images from both methods, the images are processed to generate a gene-expression matrix followed by segmentation to obtain a cell-level matrix usually via computational approaches. So let’s buckle up for some reviews of these bioinformatic analysis methods in a future post, where we can see the behind the screens of the latest techniques.
If you do not have a target in mind or you are profiling changes in spatial transcriptome under certain conditions and you are likely to take an NGS-based approach to spatial-omics, let’s briefly read through what that entails.
Next-generation sequencing-based approaches
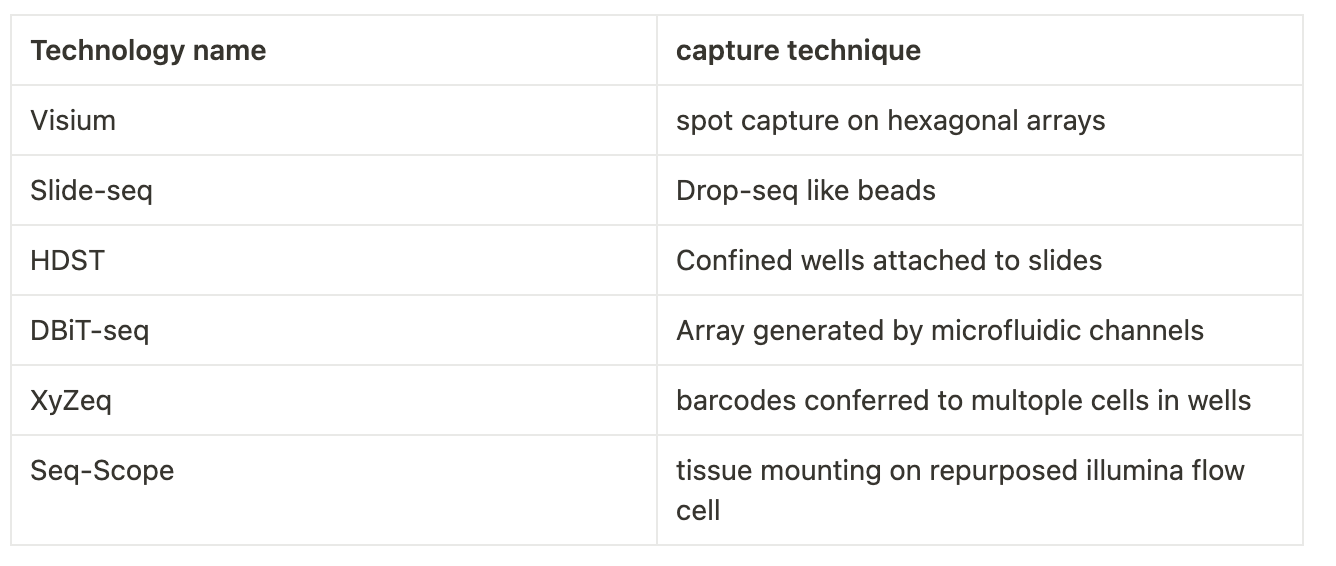
By 2016 improvements in scRNA-seq methodologies led to the first report on an NGS-based method for spatial transcriptomic that enabled the capture of whole transcriptomes from tissue sections. For this leap, a key innovation was the capture of poly-adenylated RNA on a spatially barcoded microarray slide before reverse transcription; This ensures that each transcript can be mapped back to its original spot using the position of the molecular barcode. Using this method, it was then possible to investigate larger sections of tissues in an unbiased manner. This is another cool example of technology convergence; Microarrays (hot tech of the 90s), Highthrouput sequencing(hot tech of 2000’s) and single-cell sequencing. Other NGS-based approaches.
So as in imaging based technologies, Bioinformatic analysis plays a crucial role in decoding the nature of transcript, their relative abundances and overall interpret the data into a meaningful insight into the biological question at hand. The data processing steps can be classified into upstream analysis; wherein the bioinformatician converts raw data into forms more amenable to biological interpretation and is dependent on the data-collection technology. For example smFISH- and ISS-based methods, the raw data consist of images of fluorescent spots, which must be processed to identify transcript spots, match spots to genes, and assign spots to cells. In NGS based methods a deconvolving step involving models such as negative binomial and non negative least squares are used to identify the cell type identity.
Overall we can see massive improvements in technologies available for spatial transcriptomic. Here we just had a brief introduction as a part of a multi-part series on spatial transcriptomics; next week lets dive into one technology and look through a research paper that has either introduced that technology or that has leveraged the technology to gain new insights. I hope this has been informative; Please spread the word around to your friends and colleagues about the blog. I very much appreciate giving me a chance to share my interests with such lovely curious people.
References and further reading.
Museum of spatial transcriptomics by lambda Moses and Lior pachter
Spatial omics technologies at multimodal and single cell/subcellular level by Jiwoon park and co-authors
Exploring tissue architecture using spatial transcriptomics by Anjali Rao and co-authors
Do you like the article? And do you want a place to hang out, talk, and be the nerd you are meant to be? Subscribe now for free and spread the word!!!
oh, If by you gained some insights and would like to help me be caffeinated to write the posts please scan the QR below or click here :)