Tech Focus: Molecular Pixelation
A new frontier in single-cell spatial proteomics
Dear Valued Community Members,
Navigating the complex world of scientific research and translating it into understandable terms for the general public is a challenge I'm eager to embrace. Deciphering jargon-laden papers and transforming scientific knowledge into digestible nuggets of information is no easy feat, but I'm committed to making science accessible to all.
To achieve this, I'm experimenting with different writing styles, exploring diverse techniques, and even employing a touch of humor to make science engaging and enjoyable. Remember, even the most serious subjects benefit from a dash of levity.
Join me on this journey of scientific translation as we bridge the gap between science and society. Let's make knowledge accessible to all, not just a select few. Stay tuned for more science-infused adventures, and don't hesitate to share your thoughts and suggestions along the way. If you find the post helpful, it would be great if you could leave a like or a comment. And if you really love it, please feel free to share it with your friends on Twitter or Bluesky.
Happy reading!
Our bodies are incredibly complex, and the similarity is striking when we compare them to everyday objects or places. They can be likened to a metropolis of modern times, with a multitude of cells constantly on the move, navigating through the highways of blood vessels, playing a vital role as an agent of infrastructure, supporting the movement of cells and their constituent information.
However, it's unfortunate that many individuals who are not part of the field of science tend to ignore this complexity. The nature of this complexity is often overlooked and not pondered upon. It's essential to appreciate and understand the intricacy of our bodily functions and how evolution has led us to this point.
Furthermore, the cells themselves contain a bustling metropolis of molecular activity. The symphony of proteins that orchestrate these activities carry out the essential tasks that define life, from building tissues and transporting signals and nutrients to catalysing chemical reactions and enabling communication between cells. Our bodies are a true marvel of nature, and it's essential to appreciate the intricate workings that make them so extraordinary.
Researchers have traditionally relied on immunofluorescence methods to gain insights into complex cellular processes. These methods involve using specialised antibodies to illuminate particular proteins on cell surfaces and determine their relationship with neighbouring cells. Additionally, these methods can be used with other techniques, such as microscopy, flow cytometry, and Fluorescence Resonance Energy Transfer (FRET), to understand cellular function better.
Molecular Pixellation: Single-cell spatial proteomics by sequencing.
Molecular Pixelation represents a ground-breaking approach that harnesses the power of DNA sequencing to analyse individual cells and measure the amount, distribution, and overlap of specific proteins using Antibody Oligonucleotide Conjugates (AOCs). By grouping AOCs into local clusters based on their relative positions, using DNA pixels with unique identifiers, this method enables the generation of spatial maps of individual cells. With each cell divided into over 1000 interconnected spatial zones in three dimensions, this technique offers an inspiring new way to explore the complexities of life. The DNA-sequencing reads are then computationally arranged to create maps of 76 proteins without cell compartmentalisation, opening up new possibilities for the future of scientific research.
Key Idea behind the technology.
MPX utilises DNA-tagged antibodies bound to protein targets on chemically fixed cells to conduct a comprehensive and multiplex analysis of protein surfaces.
Unlike 10x, our assay can be performed in solution without needing compartmentalisation of cells.
It also uses a unique molecular identifier to detect the spatial arrangement of proteins by serially forming associations between spatially proximate AOCs in local neighbourhoods via UMI.

How does MPX work?
The demonstration provided a fascinating insight into the spatial organisation of immune cell surface receptors in a three-dimensional space. DNA pixels, each with a Unique Pixel Identifier (UPI), enabled them to create neighbourhoods of AOC molecules with identical UPI sequences through hybridisation and gap-fill ligation. The second set of DNA pixels, incorporated through hybridisation and gap-fill ligation, was amplified by PCR and sequenced, and the resulting molecules contained a unique plate number, a protein barcode, and a barcode with neighbourhood memberships. To locate each AOC molecule, they employed computational modelling, which allowed the creation of a bipartite graph where unique molecules were represented as edges, and UPI-A and UPI-B sequences were nodes, with the protein identity serving as edge attributes. They also considered an alternative approach of a one-mode projected graph where UPI-A sequences were nodes, and protein identities were edges. The demonstration showcased an innovative and exciting approach to studying the spatial organisation of immune cell surface receptors, which could pave the way for future research in this field.
Bipartite graphs and what now...
I understand that interpreting domain-specific terminologies can be difficult, so here is a simplified version to help you know.
Imagine you have a box of crayons and a blank sheet of paper in front of you. You can draw anything you want on that paper using those crayons. Now, think of those crayons as the vertices of a graph and the lines you draw as the edges of that graph. Exciting, isn't it?
Let's take it up a notch and give it a fun twist. Imagine you want to draw a picture of a cat and a mouse. You can use your favourite colours to draw different parts of the cat and mouse. For instance, you could use blue crayons for the cat's eyes and yellow ones for the mouse's whiskers. That's super cool, right?
Now, here's where it gets even more enjoyable. Edge attributes are like the different colours of crayons you use to draw those cute little creatures. Just as different colours can tell you other things about a picture, edge attributes can tell you different things about a graph. For example, an edge attribute could tell you how strong the relationship is between two vertices.
In the cat and mouse example, the edge attribute could be the colour of the line you draw between them. If you draw a red line between them, it could mean that the cat is chasing the mouse. But drawing a green line between them could mean that the cat is friendly with the mouse. How cool is that?
That's what edge attributes are all about. They add more information to a graph and better help you understand the relationships between vertices. So, next time you draw something with crayons, think of it as a fun little graph and go wild with those edge attributes!*
Demonstration using Peripheral blood mononuclear cells.
The donor cells were fixed using Paraformaldehyde, which cross-links proteins and preserves cellular morphology for subsequent analysis. The assays for this demonstration were performed in replicates. The first replicate was subjected to staining and performing MPX, while the second was used as a PCR-based library for sequencing.
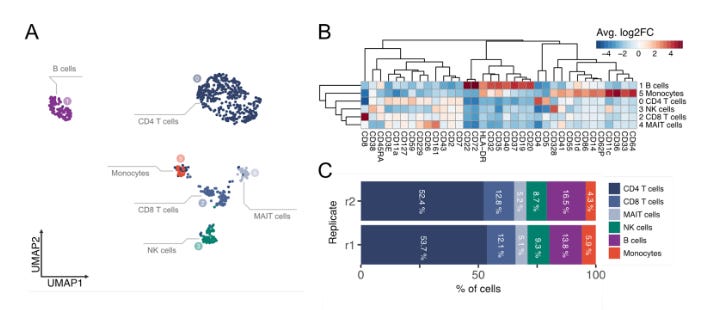
The protein count data associated with MPX was processed to generate clusters using UMAP. The results show that the clusters separated well and corresponded to the various cell types such as T cells, B cells, NK cells, and monocytes. The cell types were inferred from the differential abundance of proteins.
Demonstration of colocalising protein pairs in chemotactic T-cells
T cells are a type of white blood cell that help our body fight off infections and diseases. When we get sick, T cells need to move towards the site of infection to help fight it off. To do this, T cells follow chemical signals called chemokines that act like breadcrumbs, leading them to where they need to go.
Chemotaxis is the process by which T cells move towards these chemical signals. It involves a complex series of events inside the T cell, including activating specific molecules and rearranging the cell's structure.
Some crucial parts of the T cell that help with chemotaxis include:
1. Chemokine receptors: These are like sensors on the T cell's surface that can detect the presence and direction of the chemokine signals.
2. Integrins are like anchors that help the T cell attach and detach from the surfaces it moves across.
3. G proteins are like messengers that help relay the chemokine signal from the receptor on the T cell to the cell's internal machinery that drives its movement.
Chemotaxis plays a crucial role in the functioning of T cells by aiding them in locating and arriving at the site of infection, where they can battle the pathogens responsible for the ailment. Moreover, it assists them in moving to various parts of the body as required, such as lymph nodes, where they can be stored as "memory" cells to combat future infections.
Using MPX and a colocalisation score based on the Pearson correlation coefficient and permutation testing, scientists can estimate the degree of subcellular spatial co-occurrence of two proteins. To demonstrate how MPX is applicable in the real world, they showcase its ability to detect cellular structures called Uropods.
The uropod is a structure of migrating T cells made up of actin filaments and myosin motors. It plays an important role in T cell chemotaxis by providing adhesion and traction during migration. Its formation is triggered by chemokine receptors on the T cell surface, leading to the polarisation of the cell and the formation of a leading edge. The actin filaments in the uropod are anchored to the cell membrane, providing a platform for the contractile activity of myosin motors.
The uropod is crucial in various cellular processes, including T cell chemotaxis. It helps T cells adhere to endothelial cells and generate the force required to cross the endothelial barrier during transendothelial migration. Additionally, the uropod aids in stabilizing T cell position during synapse formation with antigen-presenting cells and maintaining contact with them.
In the lab, scientists used different methods to make cells migrate and form uropods. They tested how adding certain chemicals affected the process and compared it to a control group that wasn't stimulated at all. They measured protein levels, polarization scores, and how much different proteins were overlapping with each other. The most exciting results came from the cells that were immobilized by one of the methods and then treated with certain chemicals. They found 12 proteins that were significantly different and 15 markers that were arranged differently. Some of the markers, like CD2, CD44, and CD50, had a different pattern when the cells were immobilized and stimulated. But the most exciting part was that they found increased overlap between CD162 and CD50, which could have important implications for future research.
Overall, these demonstrations show that MPX can be employed to identify patterns of protein spatial organization and and their potential roles in cellular processes, Identificantion of markers that are associated with cell surfaces, and for studies on cellular signalling.
Some afterthoughts.
The complex network of proteins on the surface of cells and their interactions with neighboring cells could provide vital clues to a wide range of diseases, even rare ones like Epidemolysis bullosa caused by genetic mutations affecting cell adhesion proteins. By utilizing this platform, not only can we gain deeper insights into these diseases, but also monitor and improve health outcomes. This platform can also aid in refining our understanding of pathogens, such as giardia, by studying its impressive movement mechanism, or help us understand viral entry by identifying the proteins on the cell surface that are activated by viral particles, thereby enabling the development of specific therapeutics.
References:
Molecular Pixelation: Single cell spatial proteomics by sequencing
Molecular Pixelation is a commercially available proteomics analysis kit and is sold by Pixelgen Technologies AB. Read more on their site here or visit their LinkedIn page
All images are obtained from their research article available on biorxiv.
From the Tech focus series:
Tech focus: Akoya Phenocycler-Fusion
Dear Members of the community, I'm confident that you've come across numerous articles about spatial biology and its potential applications, including some from me and other experts. You may have noticed that my previous posts have primarily focused on this topic. My goal is to help you - whether you're a researcher or simply someone curious about the fi…
Peering through the tissues #6: The next frontier
Dear Members of the community, I hope you are having a well-deserved easter break, I am going to keep this short for the sake that you can enjoy the time with your family and loved ones. It has been a thoroughly enjoyable experience reading the paper that I try to summarize today. It is exciting to see that spatial transcriptomics as a sub-field of biolo…
Peering through the tissues #4: Iterations
Dear Community, This week’s post will likely be rather short. I am heading to the Netherlands to meet a buddy over the weekend. As much as I want to write a detailed breakdown of two cool papers, I have to cut it short, so the format will be slightly different. But I am up for a chat in the subscriber’s chat to clear your doubts and satiate your curiosit…
If you found the article valuable or if it brought a smile to your face, it would mean the world to me if you could take a moment to leave a like and a comment below. Moreover, if you are feeling generous and would like to support my writing endeavours, you can scan the QR code to make a donation and perhaps buy me a cup of coffee. I genuinely appreciate your thoughtful consideration.