Dear Members of the community,
I'm confident that you've come across numerous articles about spatial biology and its potential applications, including some from me and other experts. You may have noticed that my previous posts have primarily focused on this topic. My goal is to help you - whether you're a researcher or simply someone curious about the field - better understand the available methods and technologies in spatial biology. As such, I'm starting a new sub-series that showcases various technology demonstrations in spatial biology. I recently had a moment of realization when I compared the number of publicly listed spatial biology companies (thanks to
) to my circle of competence (a term commonly used among investors). It's apparent that the biotech industry, particularly market leaders in spatial biology, is significantly undervalued. Through these tech-focused articles, I aim to provide a more comprehensive understanding of the current market trends, limitations, and opportunities in spatial biology.Dupin of the Unseen
In the realm of the unseen exists a hidden world of cells and molecules performing a symphony of life. This world has long been shrouded in mystery, but with the emergence of spatial biology, we can finally observe its depths.
The Akoya Phenocycler-Fusion system is an impressive toolkit, a machine that enables us to map the spatial distribution of cells and molecules in tissues with unparalleled precision. It is a technology that has the potential to revolutionise our understanding of biology and disease.
Imagine a world where we can detect the cancer cells hiding within a tumor, or the immune cells rushing to fight infection. Imagine a world where we can map the neural circuits responsible for brain function or the molecular pathways that lead to disease development or deconvolute cells involved in complex cancers.
Spatial biology promises this world. A world where we can finally see the unseen and where this new knowledge can be utilised to improve human health. In the hands of a proficient researcher, the Phenocycler-Fusion system is a powerful tool for discovery. It can be used to answer questions that have long perplexed scientists and reveal new insights into the workings of life.
For instance, spatial biology has allowed researchers to identify new cancer biomarkers and therapeutic targets. It has also been used to study the immune system in unprecedented detail and to map the neural circuits that underlie brain function.
Spatial biology is rapidly advancing, and discoveries are regularly made. As the technology continues to develop, we can expect to see even more groundbreaking insights into the biology of health and disease.
The Akoya Phenocycler-Fusion system is a powerful tool that aids us in exploring the unseen world of cells and molecules. It is a technology that has the potential to revolutionise our understanding of biology and disease and to help us create a healthier future for all.
Mapping the Spatial Proteome of Head and Neck Tumors: Key Immune Mediators and Metabolic Determinants in the Tumor Microenvironment
HNSCCs, also known as head and neck squamous cell carcinomas, are tumours that develop in different regions of the face and neck, including the lip, oral cavity, and larynx. Even though this group of cancers is well studied and immunotherapies are available, there are currently no validated biomarkers to predict the responsiveness to immunotherapy across all HNSCC due to a significant degree of intra and inter-tumoral heterogeneity.
Spatially contextual biomarkers can provide valuable guidance in the analysis of cancer. By examining the constituent cell types of the tumour microenvironment, we can gain insight into the cellular makeup of cancer. In a recent study, researchers utilised the Akoya Biosciences CODEX platform (renamed Phenocycler™)to evaluate a comprehensive 101-plex antibody panel. This panel included biomarkers related to immune checkpoints, tumour-promoting inflammatory biomarkers, angiogenesis, invasion, and metastasis and pathways associated with cellular energetics, proliferation, evading growth suppression, and apoptosis.
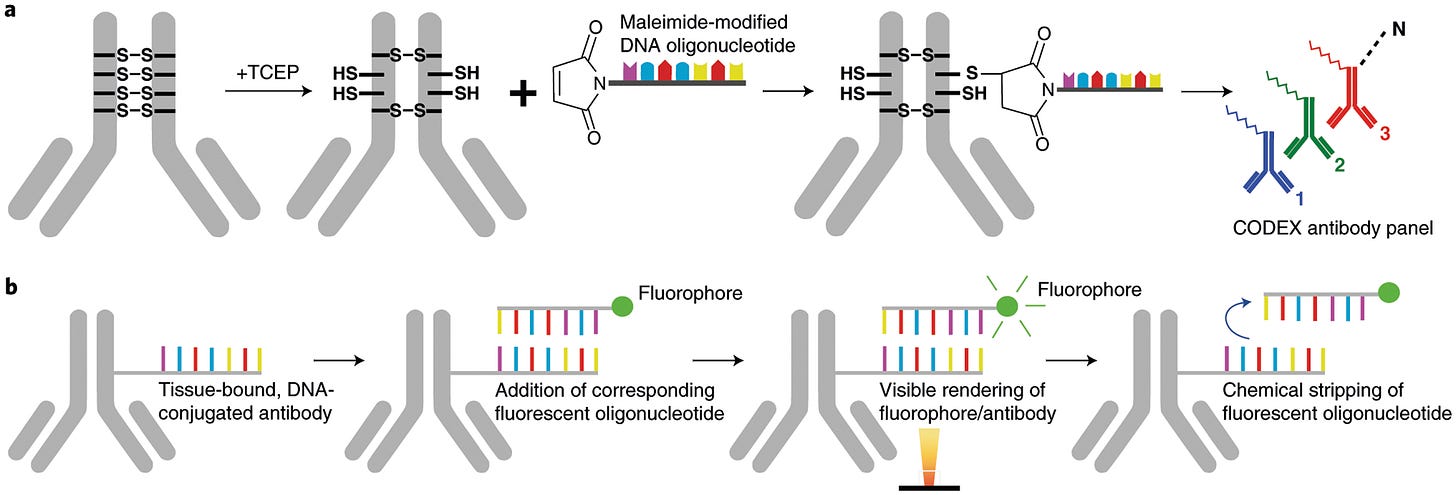
Process overview of the platform
Fluorescence microscopy platform based on the detection of DNA-conjugated antibodies.
It uses an automated microfluidics system and a conventional fluorescent microscope to iteratively hybridise, image, and strip fluorescently labelled DNA probes that are complementary to the tissue-bund DNA conjugated antibodies.
Enables single staining procedure, short run times simple chemistry and simultaneous visualisation and quantification of up to 60-101 markers in a single tissue section.

101-markers for the Human tonsil and HNSCC.
The study discussed the use of high-plex single-cell spatial phenotyping to understand the human Tumor microenvironment (TME). The authors developed a panel of markers that could identify various aspects such as cellular metabolism, apoptosis, tumour invasion, metastasis, and cellular proliferation. The performance of these antibodies was evaluated by comparing their staining patterns with publicly available data sets, followed by optimisation for maximum signal-to-noise ratio. The reproducibility of the results was assessed by running the panel on the tonsil and tumour tissue. On the technical side, the spatial localisation of all 101 proteins across the entire tissue section was revealed by imaging three markers per cycle with a total of 38 cycles of reporter hybridisation, imaging, and de-hybridization.

The authors of the study utilised Stardist for a single-cell spatial organisation analysis of the tonsil tissue. With this method, they were able to segment the nuclei and perform an unsupervised clustering, which uncovered 11 phenotypic clusters. The researchers then assigned cellular identities based on the expression profiles of the 34 markers in each segmented cell.



Cells in tonsil tissue were spatially mapped using xy coordinates, generating a phenotypic map of cellular organisation at single-cell resolution. Correlations between immune, cell lineage, metabolic, proliferation, stress, and structural markers were explored. Co-expression of markers in certain cell types resulted in distinct expression matrices indicating the type and state of a particular cell. They give an example of ZAP70, a tyrosine kinase involved in lymphocyte activation, which colocalises with T cell lineage markers during development. Using this framework of 34 immune and structural biomarkers, they were able to annotate 14 cell phenotypes from the HNSCC tissue and were able to show the geographical distribution of all the cell types at a single-cell level. A deeper dive into this distribution was performed in the context of cellular neighbourhoods since tissues arrange themselves in complex compartments, and the following table is a broad overview of the cellular neighbourhoods and the major cell constituents. This analysis shows that the spatial distribution of the cellular neighbourhoods correlates well with previously demonstrated phenotyping data.
The metabolic heterogeneity of HSNCC.
In this article, the authors delve into the potential of metabolic profiling in monitoring immunotherapy responsiveness as compared to CNs, presenting a detailed study on HNSCC. The study entailed analysing clusters on 23 markers involved in different cellular activities such as metabolism, invasion, apoptosis, cell cycle, proliferation, and survival. The researchers utilised an approach similar to scMEP (single-cell metabolic regulome profiling) to explore metabolic responses along with proliferation and stress pathways across HNSCC. The findings revealed a more nuanced profile across cell types, with the help of metabolic and functional features, adding an additional layer of complexity to the spatial map. Overall, the study highlights the significance of metabolic profiling in providing a dynamic snapshot of cellular activities.
Key Takeaways
One of the first studies using 101-plex barcode-conjugated antibody based spatial profiling of tissues via the optical imaging approach.
Developing these ultrahigh-plex panels are usually challenging and its a fine balance between imaging quality, antibody staining patterns, and extracting good information with low noise.
Cellular Neighbourhoods and metabolic insights in HNSCC shows a potential approach to implement new methods of disease analysis.
Large antibody panels are usually not cheap and likely a hindering factor for small-scale labs.
Spatial-Biology and its methods are in a very nascent stage.
References
Disclaimer: All the images above are from the academic paper.
If you found the article valuable or if it brought a smile to your face, it would mean the world to me if you could take a moment to leave a like and a comment below. Moreover, if you are feeling generous and would like to support my writing endeavors, you can scan the QR code to make a donation, and perhaps buy me a cup of coffee. I genuinely appreciate your thoughtful consideration.











