Peering through the tissues #5: Layers of information
In investigatione pro magis notitia: into the DBiT-Seq universe
Dear Members of the community,
Sorry for the delay in delivering the newsletter, I had to work on my PhD project which took a significant portion of time as I am trying to wrap it up.
The last few weeks have been an exciting time for spatial transcriptomics, there have been several new and exciting papers leveraging spatial-omics platforms. Since I started writing early this year, I am under the impression that there is a significant adoption of spatial transcriptomics. Especially for understanding basic biological processes where there was a missing layer of information, ie how do cells and their transcriptomes communicate and form the organisational level of the tissue?
The point at which we are seeing rapid progress in new experimental and computational methods in spatial transcriptomics is where sc-SEQ was a few years ago. At this juncture, it is becoming slightly challenging to keep up and write about all the tools and methodologies that are being published. But I will try my best to keep you all informed, I guess I will be adding a few papers as an additional resource for your essential reads and if someone requests a long-form summary I can write it up.
As always, take the action to subscribe now. Hit that button and share it with your friends and colleagues
Nothing has such power to broaden the mind as the ability to investigate systematically and truly all that comes under thy observation in life.
— Marcus Aurelius
Genetic information in the Eukaryotic cell is organized in a highly complex manner, I would also argue that evolution has not yet perfected this organization and yet we come across a really fine-tuned machinery which interacts with the levels of information. As a layperson, terms such as DNA, RNA and protein are most widely known and recognized as some sort of information mechanisms crucial for the survival of the organism. But I would argue that we should rephrase this information hierarchy into much more elegant categories that would give us more pliancy to introduce a wider context of genetic information. One proposal that is quite more common is the arrangement of genetic information into a sequence, gene order, gene family and epigenetics/spatial structure of the information.
At the lowest level, the sequence is composed of building blocks of nucleotides in the form of DNA. These at a first glance appear to be random, but they are far from it. The DNA is arranged into a sequence of genes which are then packed tightly by proteins called Histones. It is estimated that the total extension length of all the nuclear DNA molecules presents in a single human individual is appx 6.2 billion kilometres; To put it into the context of the solar system, it is 1.05 times the distance between the Sun and Pluto. With this perspective, you can see how efficiently the DNA is packaged and arranged for “cellular life activities”.

Heritable traits are often associated with changes in the DNA itself, the most popular familial diseases are associated with heritable changes in the DNA. In a similar fashion, epigenetic changes also play a crucial role in the passage of information changes through time. Simply put, Epigenetics is the study of heritable changes in gene expression that are not caused by changes in the DNA sequence itself. These changes can occur through modifications to the DNA molecule, such as methylation or acetylation, or through modifications to the proteins that package and regulate DNA, such as histones. Epigenetic modifications are important for regulating gene expression during development and differentiation and can have long-lasting effects on an organism's phenotype. For example, during embryonic development, epigenetic changes help to activate or silence genes in specific patterns that determine the formation of different tissues and organs.
For this week’s dive into a new method (spatial epigenome sequencing), I have to re-wind a bit and dwell on previous methods that are crucial and contributed directly to the development of the current method.
1. ATAC-SEQ:
Assay for Transposase-Accessible Chromatin using sequencing. A method developed by Jason D. Beunrostro and collaborators. ATAC-Seq is an essential molecular biology technique used to analyze chromatin accessibility. Chromatin refers to the combination of DNA and proteins that make up chromosomes within a cell. The accessibility of chromatin can affect gene expression, and ATAC-Seq allows researchers to identify regions of DNA that are open or accessible for regulatory proteins to bind to and activate or repress genes.
ATAC-seq captures open chromatin sites using a simple 2-step protocol from 500 to 50,000 cells (as originally described), and reveals the interplay between genomic locations of open chromatin, DNA binding proteins, individual nucleosomes, and higher-order compaction at regulatory regions with nucleotide resolution. But the single-cell versions have become more of the norm.
How does it work?
Transposons are genetic elements that can “jump” to different locations within a genome. The first transposons were described by the legendary Barbara McClintock in her studies on maize. ATAC-Seq frequently uses Tn5 transposon’s enzyme the transposases that were discovered in Escherichia coli consist of a core sequence encoding three antibiotics (neomycin, bleomycin, and streptomycin) and two inverted IS50 sequences, IS50L and IS50R, which encode a Tn5 transposase (Tnp). Essentially, this in the bacteria provides a toolset to fight antibiotics and excision followed by insertion “jumping” can provide resistance against the antibiotics for a bacterial colony.
In genomics research, The “cut and paste” function of Tn5 is widely used. Studies have shown that only Open End sequences and Tn5 transposases are required for transposition in vitro. Tn5 transposases can randomly insert adaptors/barcodes into DNA, and the resulting DNA molecules are ready for PCR amplification and sequencing.
The following is a video that beautifully summarizes ATAC-SEQ
2. CUT and TAG sequencing:
Cleavage Under Targets and Tagmentation (CUT&Tag) is an enzyme-tethering strategy that provides efficient high-resolution sequencing libraries for profiling diverse chromatin components. The method was described by Hatice S. Kaya-Okur and collaborators. The method similar to ATAC-seq relies on Tn5 transposase with some variation in the strategy.
CUT&Tag leverages the use of antibodies which bind to target chromatin protien between the nucleosomes in the genome. This is followed by a secondary antibody that allows the tethering of protein A-Tn5 transposomes. This is followed by an activation step where the transposomes are activated and the integration of adapter sequences at the chromatin protein binding sites.

Below, is a video explaining CUT&Tag:
Spatial epigenome-transcriptome co-profiling of mammalian tissues.
Having given a brief overview of two technologies that played a crucial role in our understanding of the epigenetic architecture of the cells in disease and development. We can dive a bit further into the integration of these technologies along with DiBT-seq to co-profile the transcriptome and the epigenome in a spatial context.
I have previously written about Deterministic barcoding in tissue for spatial omics sequencing (DBiT-seq) in the post from the 4th of march, in the context of Spatial CITE-Seq. In this current long-form summary let’s see how the teams from Yale led by Prof Rong Fan, and Karolinska Institute led by Prof Gonçalo Castelo-Branco have delivered us with an exciting new method that allows us to jointly profile the epigenome and transcriptome.
This is important because spatial profiling till now has been limited to a single snap-shot of a particular information layer. It limits our ability to understand what is happening at the other layers of the information hierarchy. For example, in Epstein Barr Virus positive cancers, it is known that LMP1 a latent membrane protein produced by the virus selectively hypermethylates tumour suppressor genes leading to the promotion of tumour growth. But this is not the only thing that seems to happen in such infections, There are broad changes that suppress the host transcription and translational machinery for which classical experiments have only provided a static snapshot on each layer. By this I mean, it was likely only feasible to measure the changes at one level, such as the RNA or the Epigenome in a cell sample or tissue.
Although computational methods have been useful in integrating the multiple omics data types, they can not uncover links between different layers at a single time point concurrently. In the current paper, the authors have proposed and developed spatial-ATAC-RNA-seq and spatial CUT&Tag in the following manner.
The proposed methods use the DBiT-seq platform. Check my previous post here.
As a researcher, you have the option to profile the chromatin accessibility and mRNA expression using ATAC-RNA-seq x DBiT-seq aka spatial-ATAC-RNA-seq
histone modifications and mRNA expression using CUT&Tag-RNA-seq x DBiT-seq aka spatial CUT&Tag-RNA-seq.
They tested and validated the methods on embryonic and juvenile mouse brains as well as adult human hippocampus to dissect the roles of epigenetic and transcriptional states and how is their interplay in the regulation of cell types in the tissue.
Methods of spatial epigenome-transcriptome profiling
For spatial ATAC-seq:
the authors used a frozen section of tissue that was fixed with formaldehyde and treated with a Tn5 transposition complex preloaded with DNA adapters consisting a universal ligation linker similar to what was described above.
The consequent step is the incubation of a biotinylated DNA adaptor (a modified DNA sequence that can be treated with streptavidin later to capture it). This also contains special sequences that allow it to bind to the targets of interest ie mRNA to initiate the reverse transcription.
In the fashion of DBiT-Seq, a microfluidic chamber chip was then placed onto the tissue to introduce the spatial barcodes. The spatial barcodes bind to one of the special sequences via templated ligation.
This is followed by a second set of spatial barcodes resulting in a two-dimensional grid of spatially barcoded tissue pixels each having a unique combination of barcodes as discussed in the previous post.
cDNAs can be enriched with streptavidin beads and genomic DNA fragments are present in the supernatant. Here the cDNA component represents the transcriptome and the gDNA is the accessible component of the chromatin.
For spatial CUT&Tag-RNA-seq:
They used antibodies specific against histone modifications to the tissue section in a similar fashion to the normal CUT&Tag mentioned above.
Using a protein A-tethered Tn5-DNA complex they performed CUT&Tag, with the remaining size similar to spatial-ATAC-seq
Data Quality
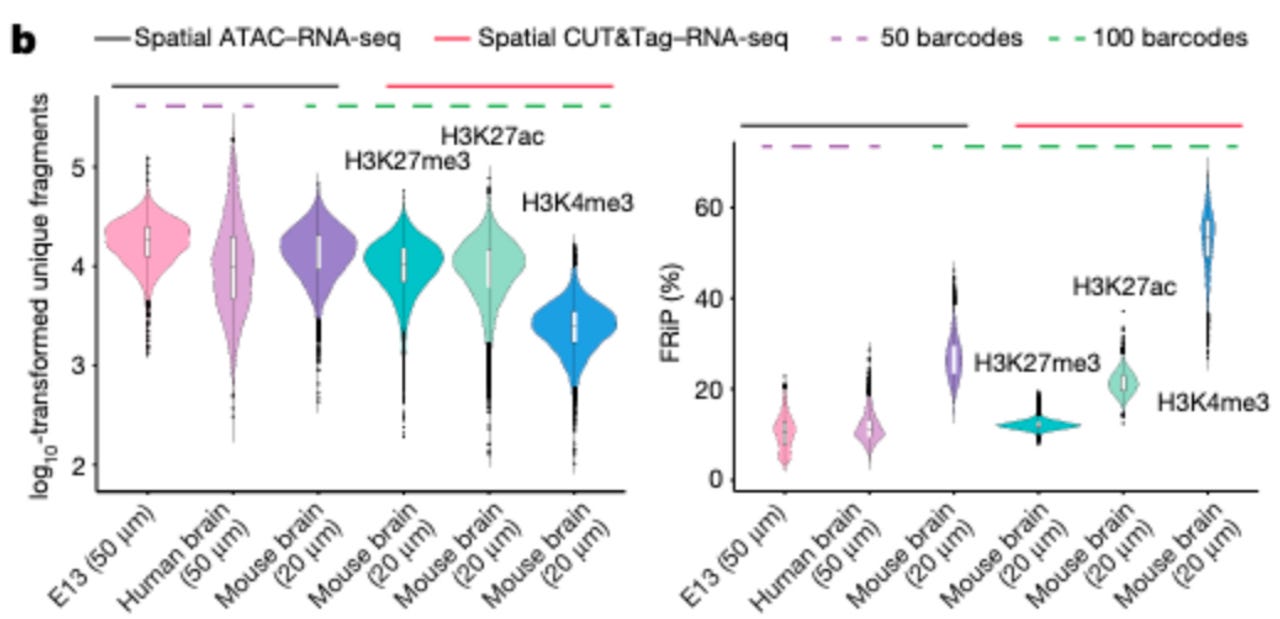

Key takeaways from spatial co-mapping of the mouse embryo.
Spatial ATAC-RNA-Seq of E13 embryo identifies 8 major ATAC clusters and 14 RNA clusters
From ATAC data, cluster A3 represents the embryonic eye. A4-A5 are associated with the development of internal organs. A6-A7 cover the CNS
They performed benchmarking of ATAC using organ-specific ENCODE E13.5 ATAC-seq. They also integrated this with single-cell gene expression.
In the brain, they were expecting to see neural stem cells predominantly present in the ventricular layer. And much more mature cells were distant from the ventricular layer.
Genes such as Sox2 which is involved in the development of nervous tissue and optic nerve showed high chromatin accessibility in the eye region and ventricular layer.
Six6 a gene involved in eye development showed the highest Gene Activation Score (GAS) in the eye region. Indicating that this method is sensitive to measure chromatin accessibility while retaining spatial information.
For the RNA part of the spatial data, the 14 different clusters were characterized by specific marker genes.
Like the above cluster, R10 was identified to be correlated to the embryonic eye by measuring the Six6 gene.
Similarly, they see a few more clusters correlate to organ-specific RNAseq reference data. Indicating that there is a high concordance between the data generated and the reference dataset.
They were also able to identify a few cell type-specific markers like Mapt in cluster R2 that might play a role in establishing and maintaining neuronal polarity.
By joint clustering of spatial ATAC and RNA data to refine spatial patterns, the authors were able to identify a new neuronal cluster that was not seen by using single methods.
They find that chromatin accessibility is not directly correlated to RNA expression but is more likely that the accessibility influences the distribution of RNA.
Also, increased accessibility at this particular time point is likely due to the priming of gene expression for the next stages of the organism’s progression.
By utilising a pseudo-time analysis they were able to visualize the developmental trajectories. The chromatin accessibility and gene expression along these trajectories show dynamic changes in selected markers

Above I have provided a summary of the extensive results from the paper. This is one part of a multiple-sample study where they show that this method can be applied across various tissue types. I won’t deep dive into other samples and technology types, due to the direction of the newsletter is to give you enough insights for you to get started in such cool technologies. I will go a bit deeper into the bioinformatics analysis of the data generated.
Data handling:
By using the linker sequences from ATAC and CUT&Tag the authors filtered the read 2 and these sequences were converted to Cell Ranger ATAC v.1.2 format and which consists of the newly formed genome sequences
By filtering out the genome sequences they were able to identify the barcoded sequences corresponding to the spatial transcriptome.
The human or mouse reference assemblies (GRCh38 or GRCm38) was used to obtain fastq files and BED files which were used for the downstream analysis.
ST pipeline v1.7.2 was used to map the processed reads for spatial transcriptomics.
Using Hiplex Proteome and its inbuilt functions, they identify the location of pixels on tissue from the bright field image.
ATAC, CUT&Tag and RNA matrices were analysed using Signac v.1.8
RNA: the data was normalized using the SCTransform function and subsequently the UMAP was built
ATAC/CUT&Tag: They used the DefaultAssay function followed by applying a minimum cutoff set with the FindTopFeatures function. The data were subjected to normalization and dimensionality reduction followed by clustering and projecting the clusters into the UMAP space.
For visualization of RNA spatial data gene matrix obtained was loaded to Seurat as a Seurat object and RNA metadata obtained from Signac was read into the Seurat object.
The spatial maps were generated using the SpatialPlot function.
Further, they use ArchR to read in the ATAC/CUT&Tag fragments followed by combining the metadata obtained from Signac and this is then used to normalize and reduce the dimensionality. The visualization step is performed in Seurat.
They extensively use Seurat for RNA data integration and cell type identification. Signac and Seurat for integration of ATAC/CUT&Tag and their single-cell variants (not discussed in detail in this newsletter but check the paper). Finally, ArchR was used for cell type identification for ATAC/CUT&Tag from scRNA-seq data.
Data and code availability:
Hiplex proteome: https://github.com/edicliuyang/Hiplex_proteome
Signac: https://github.com/stuart-lab/signac/releases/tag/1.8.0
Code for the profiling: https://zenodo.org/record/7395313
Raw data: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE205055
Conclusion:
As mentioned in several posts now, scientific breakthroughs occur in iterations more often than the eureka moments. Here we are presented with a method that utilizes previously available technologies and combines them in an elegant way to reveal more than one layer of information in regard to spatial tissue profiling. I can imagine that there might be multiple other teams that are working on similar approaches, maybe a spatial DamID-seq or a spatial-ATAC-seq with a single-cell resolution. I can also imagine that there are a few challenges that can be computationally addressed, one of these is the deconvolution of the results using computational approaches to achieve single-cell resolution.
This is a rapidly expanding field of study with more exciting methods that are coming into existence and also they are being applied to a plethora of challenges. I am hopeful to see new insights being gained and more complex problems being tacked using spatial transcriptomics or epigenomics.
References:
Kaya-Okur, H.S., Wu, S.J., Codomo, C.A. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun 10, 1930 (2019). https://doi.org/10.1038/s41467-019-09982-5
Zhang, D., Deng, Y., Kukanja, P. et al. Spatial epigenome–transcriptome co-profiling of mammalian tissues. Nature (2023). https://doi.org/10.1038/s41586-023-05795-1
Lab Socials:
Fun fact: I am so happy to see Dr Branco’s work evolve over time. Its been approximately 10 years since I last met him to be interviewed for a PhD student position when he was starting out at Karolinska. It is so cool that several of the methods that he has published are being widely used across biology.
If you enjoyed the article, Please leave a like and comment below. And if you appreciate the work, maybe scan the QR code and buy me a coffee.
See you next week with a cool new paper on a similar technology that is going to change the field.




