Achieving Optimal Gene Selection Accuracy with Machine Learning: The Dominance of Predictive Analytics.
Praedictio Genetica Selectiva. On PERSIST: A gene panel selection tool for spatial transcriptomics
Dear Community,
I've had a productive month personally, making good progress on my reading list. As for my PhD project, I'm currently developing a computational drug repurposing system, which has proven to be a challenging task. The dataset is sparse and difficult to work with, but I'm up to the challenge. I'm confident in my abilities to make appropriate interpretations and strategize for a better approach. Although it's time-consuming, it's all part of my PhD training, and I'm not letting it take away from my writing summaries as best as possible.
As always, take action to subscribe now. Hit that button and share it with your friends and colleagues. Leave a comment as it helps with the algorithms :)
The ultimate goal of biology is to understand how things work, and the only way to do that is to make predictions - Anonymous
For as long as humans have existed, they likely developed the practice of prediction to comprehend events in their environment. This could be based on tangible observations, like tracking the behaviour of plants and animals to anticipate changes in seasons, or more abstract methods, such as using astrology to forecast the state of human affairs. Predicting has helped us generate hypotheses and test their accuracy through experimentation. Some of these hypothesis-based approaches have gained widespread acceptance as true, with several notable examples.
Johannes Kepler (1571-1630), a German mathematician and astronomer, used his observations of the planets to develop his laws of planetary motion. These laws accurately predicted the positions of the planets in the solar system and laid the foundation for modern astronomy.
Charles Darwin (1809-1882), an English naturalist, used his observations of plants and animals to develop his theory of evolution by natural selection. This theory predicted that species would change over time in response to environmental pressures, and a vast amount of scientific evidence has since been supported.
Edward Lorenz (1917-2008), an American mathematician and meteorologist, used his study of chaos theory to predict that small differences in initial conditions could significantly impact weather patterns. This prediction led to the development of chaos theory and helped improve weather forecasting accuracy.
The remarkable imaginative capabilities of humans have led to the undeniable fact that the tools we currently employ were once prophesied in the past. Through envisioning myriad ideas, experimenting with them, and deducing predictions from them, humans have proven their ability to shape the future.
In the field of bioinformatics, predictions are crucial for making progress and understanding data. The advancements in spatial transcriptomics, single-cell sequencing, and machine learning methods have led to the development of more sophisticated predictive technologies. When it comes to selecting gene panels for tissue probing, expert opinions and literature-based panels are commonly used to determine the most relevant transcriptional signature. By analyzing millions of single cells in reference experiments, we can gain insight into the various cell taxonomies that exist.
When it comes to Spatial-transcriptomics, Fluorescence in situ hybridization methods it is now possible to integrate the localization of a transcript and gain a contextual understanding of the cell type and its role. But, Due to its experimental design, it is often not possible to probe the entire transcriptome thereby introducing a bias by selecting a pre-defined list of genes. The current paper I am summarising addresses how one can optimally define the small gene panel for such studies.
Predictive and robust gene selection for spatial transcriptomics.
The pertinent challenge in spatial transcriptomics is the lack of spatially resolved mRNA detection measurements for thousands of genes. For now, scientists rely on scRNA-seq datasets to guide the selection process. scRNA-seq data generally serve as a surrogate measure for spatial data, this in turn might introduce noise and can be limiting in the data analysis and interpretation.
Also, the selection of the gene panel should be based on the model that is being interrogated. Say as a hepatologist, you are presented with a plethora of cell types and when you are performing a spatial experiment in a region that has a variety of cell types you would likely have to design a panel based on a specific subpopulation that is of interest and also if the genes in the panel are sensitively detected irrespective of their relative abundances.
The authors of this paper propose PERSIST: PredictivE and Robust gene SelectIon for Spatial Transcriptomics (Quite creative use of acronymization).
What is PERSIST?
PERSIST is an algorithm to select genes that can serve as valuable targets in spatial transcriptomics studies. It uses scRNA-seq data and deep learning to find a small number of highly informative genes whose expression can predict the genome-wide expression profile.
What does it do?
It trains a reconstruction model with a loss function that
Accounts for noisy gene dropouts in scRNA-seq,
incorporates expert knowledge by pre-selecting or pre-filtering genes,
It scales to huge datasets using minibatch training
It quantizes gene expression levels to account for the domain shift between scRNA-seq and spatial transcriptomics.
It operates both in a supervised and unsupervised manner giving flexibility based on experimental aims.
Overall, PERSIST positions itself as a tool that a computational biologist can reach when asked which genes can give me the most valuable information based on the cell type or experimental goals. This is key for the early stages of spatial transcriptomics when commercial panels to probe the transcripts are limited. It achieves this by implementing a deep learning architecture which is explained below.
The architecture that empowers PERSIST.
Often dimensionality reduction techniques are used to select informational genes using PCA or similar reduction algorithms which often describe linear relationships. In PERSIST the authors chose to implement a non-linear model and a quantity measure which is more appropriate for scRNA-seq data, employing this PERSIST selects a discrete set of genes rather than the linear combinations.
PCA measures reconstruction quality measures using a mean squared error loss, This is often not well suited for sparse data sets such as scRNA-seq data and doesn’t account for the noisy gene dropouts. To overcome this limitation, PERSIST employs a hurdle loss function. In hurdle models, a variable is modelled using two parts, the first which is the probability of attaining value 0, and the second part is the probability of non-zero values. This is well suited for scRNA-seq where there are excess zeros (genes with “no expression” or dropouts) that are not sufficiently accounted for in more standard statistical models.
The hurdle loss used by PERSIST, therefore, involves separately predicting each gene’s expression level and whether it is actually expressed, which lets the model explicitly represent dropouts noise in its prediction.
Using Machine learning architecture PERSIST is designed to select a precise number of genes. Input genes are selected by a custom network later that enables training with stochastic gradient descent, This layer termed as binary mask layer acts as a tool to select a user-specified number of inputs within a deep learning model. The mask itself is based on Gumbel-Softmax which lets us optimize the discrete probability distributions using stochastic gradient descent. The input of the binary mask is a vector of binarized gene expression levels (ie if a gene is expressed it is 1 and 0 if not). The output is given by the element-wise product which ensures that information that flows through the model from all genes is reduced to the user-specified number.
By default, PERSIST operates in an unsupervised manner which removes the need for labels or manual annotation. But it also can be operated in a supervised manner by incorporating cell-level annotations as the model’s prediction target. This ensures that when reference cell type clusterings are not available PERSIST can still give relevant information for the required gene selection.
The authors compared PERSISt to several widely used state-of-the-art gene selection methods such as Seurat, Cell Ranger, ScanPy and GeneBasis. Although this is not apples to apple comparison since the packages with the exception of GeneBasis do perform as a tool for the selection of gene panels.
PERSIST enables more accurate scRNA-seq expression profile reconstruction.
By training PERSIST on two scRNA-seq datasets (SmartSeq v4 and 10x) they were able to collect an initial set of 10k high-variance genes and identify panels of 8-256 markers that correspond to the normal range of FISH studies.
PERSIST was able to explain more variance and outperform methods such as Seurat, Cell Ranger and GeneBasis.
Overall higher gene numbers selected for the panel give the user diminishing returns.
Since PERSIST binarizes gene expression levels during training and unlike Seurat, Cell Ranger and GeneBasis which uses raw or log expression count, it seems to be a more appropriate way to select panels.
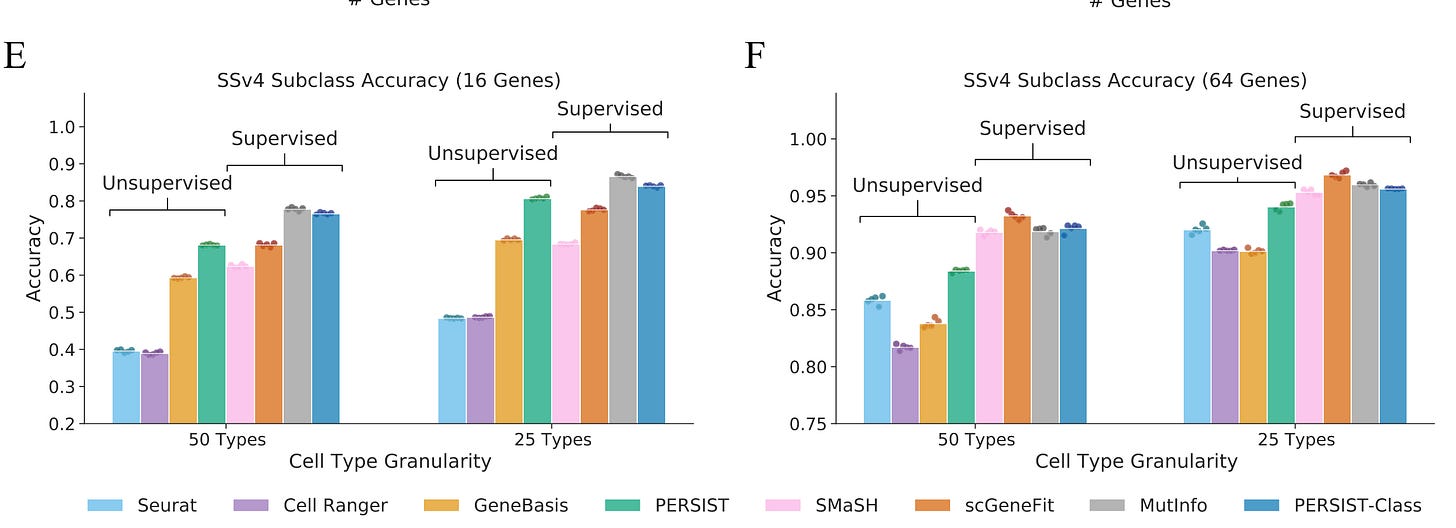
PERSIST enables accurate cell type classification
It is always challenging to identify cell types in large-scale scRNA-seq datasets. The authors test the usefulness of PERSIST on how accurately the gene panels selected by each method can classify cell types, which is somehow the goal of several transcriptomics studies. By using cell types defined by original SSv4 and 10x data, the authors show that PERSIST reaches 74% cell type classification accuracy using 64 genes in the SSv4 daya and 78% with 10x data. They term this mode of operation of PERSIST as PERSIST-Classification and it reaches around 81-82% accuracy in a supervised manner of cell classification. Furthermore, by dividing cells into broader subclasses they were able to classify cells into 25 subclasses rather than the full 113 cell types. Overall PERSIST classification provides reliably accurate cell type and subclass classifications. My personal experience with annotations of cell types in novel experimentation models is that when there is a lack of “trend” interest, no one knows what the cell types are and why they exist in samples. It is my opinion that scientists need to make bold yet arbitrary choices when it comes to labelling cells or completely misinterpret the cellular diversity, we definitely need more robust tools and agreements on the cellular hierarchies.
PERSIST can be adapted to predict electrophysiological properties.
Always, as demonstrated in previous summaries; Any new method needs to be validated to show the robustness of the method proposed. The authors developed a variant of PERSIST to identify marker genes that can predict and identify marker genes based on a multi-modal Patch-seq dataset containing transcriptomic and electrophysiological information from 3411 GABAergic neurons across 53 cell types of the mouse visual cortex. This data consists of scRNA-seq counts for 1252 curated genes and electrophysiological profiles of 44 sparse principal components and 24 biologically relevant features.
They ran PERSIST in an unsupervised manner by selecting genes that can optimally reconstruct the full expression profile followed by a supervised manner by using the electrophysiological features as prediction targets. This was then used as a model labelled PERSIST-Ephys. To evaluate PERISIT-Ephys they attempted to predict the electrophysiological features of each neuron using expression levels of the genes in each panel. Using the binarization approach described above they show that PERSIST-Ephys achieves the highest predictive accuracy for panels of various sizes.
Binarization enables gene expression prediction with MER-FISH data
Till now scRNA-seq was used to give predictive and informative markers. The challenge when it comes to FISH studies is that they are often performed in multiple panels and providing an unbiased evaluation is slightly harder when using different experiments.
For the MERFISH dataset, 258 genes were probed across 280,327 cells from the mouse primary motor cortex with no ground truth cell type labels, this forced them to evaluate performance in the expressed gene prediction task where the goal is to predict which individual genes are detected in each cell. By using SSv4 scRNA-seq datasets to select panels of 8-32 markers from within the MERFISH data. They apply an imputation model to predict which of the remaining genes are detected, this model then was used to predict the set of detected genes in the MERFISH dataset. By leveraging the Binarization of both data it is possible to transfer/match the gene expression.
Overall PERSIST reached 86.5% accuracy with a 32-gene panel. It is to be seen how exactly one can use PERSIST in real-world applications for the selection of genes and characterization of the panels etc.,
Lab socials
Su-In Lee lab: Twitter, Lab Website
Ian Covert: Twitter
References
Predictive and robust gene selection for spatial transcriptomics
If you found the article to be of value or brought a smile to your face, it would mean the world to me if you could take a moment to leave a like and a comment below. Furthermore, if you are feeling generous and would like to support my writing endeavours, you can scan the QR code to make a donation and perhaps buy me a cup of coffee. I truly appreciate your thoughtful consideration.
See you next week !!!